
RNA免疫共沉淀(RIP)
RIP(RNA Binding Protein Immunoprecipitation Assay,RNA 结合蛋白免疫沉淀)是研究细胞内RNA与蛋白结合情况的技术,是了解转录后调控网络动态过程的有力工具。RIP利用目标蛋白的抗体将相应的RNA-蛋白复合物(RBP)沉淀下来,然后经过分离纯化捕获的RNA,结合高通量测序、qPCR检测或芯片检测,对目标RNA进行分析,来鉴定结合转录RNA如mRNA、lncRNA、circRNA等,利用该技术有望发现多种RNA的调节靶点。RIP可以看成是染色质免疫沉淀ChIP技术的类似应用,但由于研究对象是RNA-蛋白复合物而不是DNA-蛋白复合物,RIP实验的优化条件与ChIP实验不太相同。
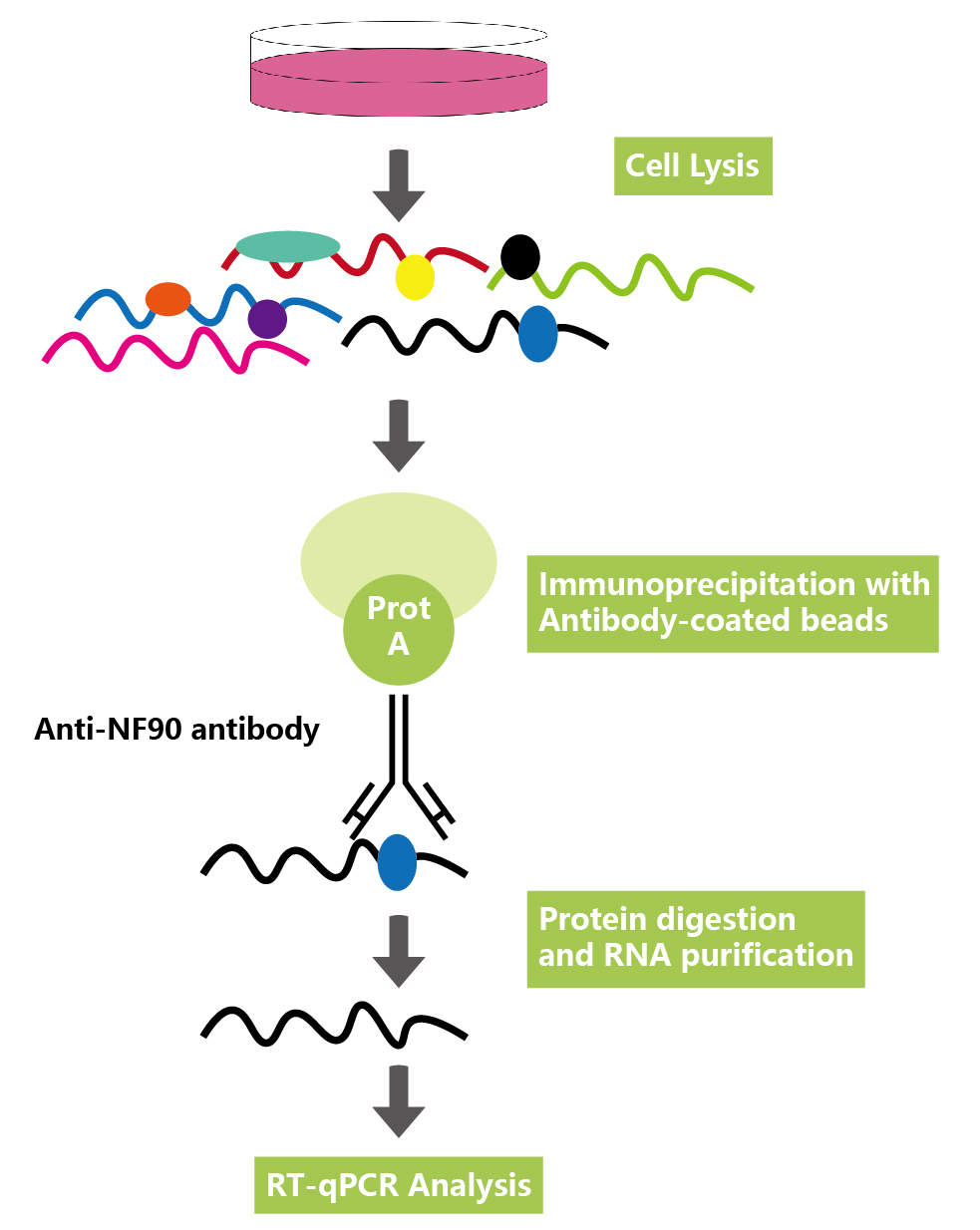
二、原理流程
RIP的基本原理是用抗体或表位标记物捕获细胞核内或细胞质中内源性的RNA结合蛋白,免疫沉淀把RNA结合蛋白及其结合的RNA一起分离出来,结合的RNA序列通过microarray(RIP-Chip),定量RT-PCR或高通量测序(RIP-Seq)方法来鉴定。在这个过程中,需要防止非特异性的RNA的结合。

三、RIP的应用
1. 研究细胞内RNA与蛋白结合情况;
2. 发现RBP与非编码RNA(LncRNA、miRNA等)的相互作用;
3. 绘制全基因组范围的RNA与RBP相互作用图谱。
案例分享:
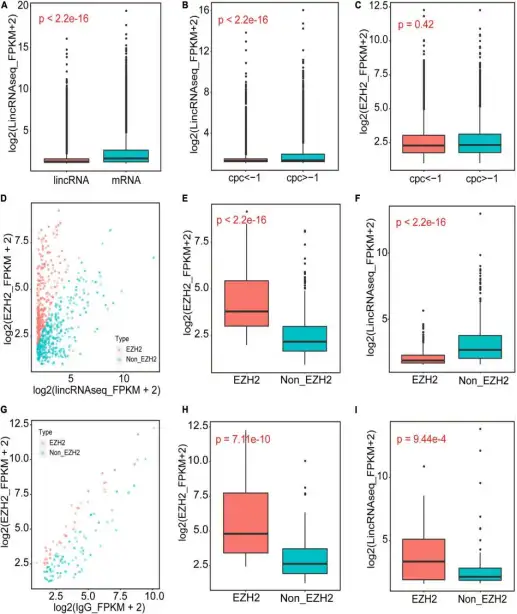
1、题目:RIP-Seq of EZH2 Identifies TCONS-00036665 as a Regulator of Myogenesis in Pigs E2H2的RIP-seq将TCONS-OOO36665鉴定为猪肌细胞生成的关键因子
研究内容:EZH2是PCR2的催化亚基,它包含一个SET结构域,可催化赖氨酸 27 (H3K27me3) 上的组蛋白H3三甲基化以产生表观遗传沉默标记。EZH2通过与转录因子或RNA转录本相互作用来实现其功能。在本研究中,作者采用RIP-seq和lncRNA-seq来鉴定EZH2结合的lncRNAs,通过生物信息学分析鉴定出356 个可以结合EZH2的lncRNA,并对其中的TCONS-00036665进行了表征。
TCONS-00036665能促进猪骨骼肌卫星细胞增殖但抑制细胞分化,这种功能在猪和小鼠之间是保守的。进一步的机制研究表明,TCONS-00036665可以与EZH2结合,并将EZH2招募到靶基因 p21、MyoG和Myh4的启动子上,从而富集H3K27me3以及抑制靶基因表达和猪肌细胞生成。
总之,lncRNA TCONS-00036665通过与 EZH2的相互作用调节猪肌细胞的生成。
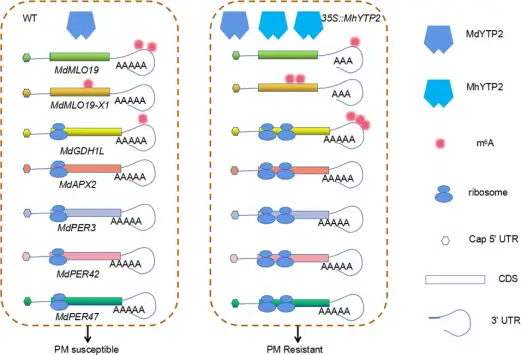
2、题目:The m6A reader MhYTP2 regulates MdMLO19 mRNA stability and antioxidant genes translation efficiency conferring powdery mildew resistance in apple m6A 阅读蛋白MhYTP2调节MdMLO19 mRNA稳定性和抗氧化基因翻译效率,从而赋予苹果白粉病抗性
研究内容:研究人员前期获得了RNA结合蛋白MhYTP2超表达转基因苹果植株,并发现转基因植株具有明显的白粉病抗性。作者首先证明了苹果的MhYTP2同样具有m6A结合功能;其次,通过检测转基因材料中m6A甲基化酶和去甲基化酶相关基因的表达水平,发现MhYTP2的超表达会改变m6A甲基化酶和去甲基化酶相关基因的表达。
借助于m6A-seq、RNA-seq和Ribo-seq等技术,研究人员发现MhYTP2过表达增加了一批mRNA的m6A修饰水平和翻译效率。同时,外显子区域的m6A修饰水平可能与mRNA的稳定性呈负相关,而非翻译区(UTR)的m6A变化与相关mRNA丰度呈正相关。此外,MhYTP2能够与含m6A修饰白粉病感病基因MdMLO19和MdMLO19-X1的mRNAs结合,并加速其降解;MhYTP2还能够结合谷氨酸脱氢酶1基因MdGDH1L的mRNA,并提高其翻译效率,增加蛋白积累,提高转基因植株的抗氧化能力。
综上所述,该研究结果揭示了苹果m6A基因谱、MhYTP2对m6A基因谱的影响以及m6A在调控苹果白粉病感病基因MdMLO19 mRNA稳定性和抗氧化相关基因翻译效率中的作用。
四、交付内容
1、原始数据、结果图片(包括阴性对照结果);
2、标准实验报告(包括详细的材料与方法);
3、SCI标准Result Figure。
五、优势特点
1、高灵敏度和高特异性:RIP技术能够高度富集RNA和与之相互作用的蛋白质,具有较高的灵敏度和特异性
2、低样本量要求:RIP技术可以在较少的细胞或组织样本中进行实验,减少实验成本和样本需求
3、适用于多种RNA种类:RIP技术适用于不同类型的RNA研究,包括长链非编码RNA和编码RNA
4、不需要事先了解RNA-蛋白质相互作用的信息:可用于探索未知的RNA-蛋白质相互作用
5、可量化分析:RIP技术可以进行定量分析,评估不同条件下的相互作用的强度和稳定性
6、结合其他技术的联合应用:RIP技术可以与其他技术相结合,如RNA测序、蛋白质组学和转录组学等,进行综合分析,深入揭示RNA-蛋白质相互作用的功能和调控机制
参考文献:
[1] Khalila AM, Guttman M, Huarte M, Garbera M, Rajd A, Morales DR, Thomas K, Pressera A, Bernstein BE, Oudenaarden AV, Regeva A, Lander ES, and Rinn JL (2009). Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. PNAS 106, 11667–72.
[2] Hendrickson DG, Hogan DJ, McCullough HL, Myers JW, Herschlag D, Ferrell JE, and Brown PO (2009).Concordant Regulation of Translation and mRNA Abundance for Hundreds of Targets of a Human microRNA.PLoS Biology 7 (11), 2643.
[3] Hendrickson DG, Hogan DJ, Herschlag D, Ferrell JE, and Brown PO (2008).Systematic Identification of mRNAs Recruited to Argonaute 2 by Specific microRNAs and Corresponding Changes in Transcript Abundance.PLoS One 3 (5), 2126.
[4] Rinn JL, Kertesz M, Wang JK, Squazzo SL, Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, and Chang HY (2007).Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs.Cell 129, 1311–1323.
[5] Martindale JL, Gorospe M, Idda ML. Ribonucleoprotein Immunoprecipitation (RIP) Analysis. Bio Protoc. 2020;10(2):e3488.
